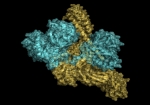
Mammalian disaggregation machinery
Protein disaggregases hold the potential to reverse protein aggregation and amyloid fibril formation – conditions that have been identified in an increasing number of debilitating, and ultimately fatal neurodegenerative disorders. In yeast, bacteria, and plants, a hexameric Hsp104/ClpB can solubilize these disordered aggregates and amyloids, however, curiously, metazoa lack an Hsp104 homolog. Instead, our cells employ a different, less understood machinery to refold protein aggregates – a chaperone system comprised of Hsp110, Hsp70, and Hsp40s.