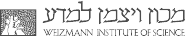Our ultimate goal is to harness our knowledge of the PCD network and its governing principles in order to more fully understand the etiology of disease in which cell death is misregulated, such as cancer.
Tumor cells carry many perturbations (mutations/epigenetic changes) along the cell death pathways, which results in cell populations that are resistant to cell death, thus limiting the efficacy of available treatments. Currently, we have several projects that investigate the integrity of the various PCD modules, and their interactivity as a whole, in various cancer models.
- By applying an siRNA library directed against each of the 81 PCD genes, and assessing the contribution of each gene to the death responses to targeted-drug therapy, we have identified the ‘functional death signature’ of individual metastatic melanoma. This information is used to rationally choose double KDs from the identified soft-spots in each tumor, predicted to enhance the initial death response to the drug, so as to reduce the probability of developing drug resistant cell populations with time.
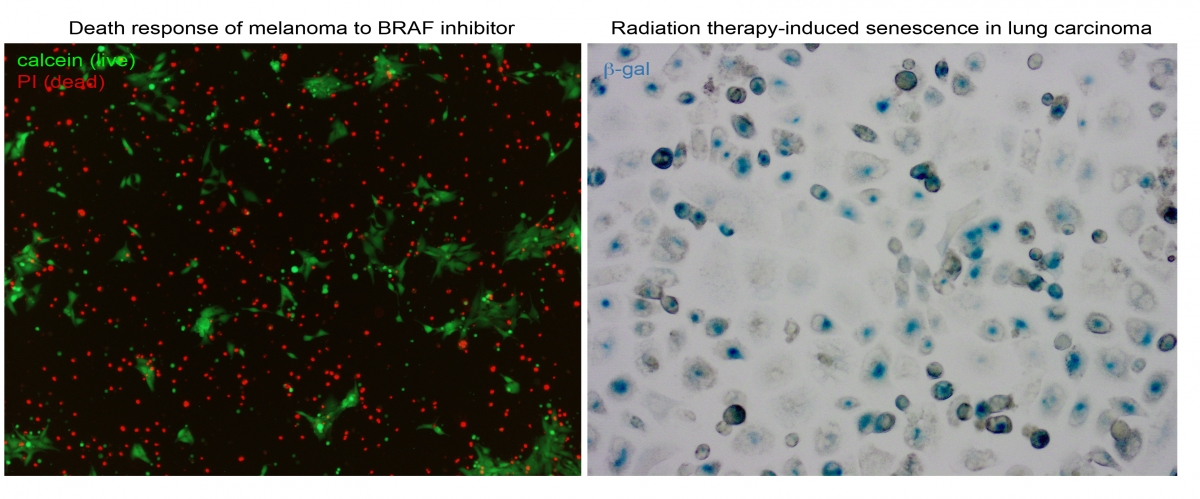
- We are examining the contribution of PCD proteins, including members of the DAPK family, to cell death pathways that are activated by radiation therapy, with the goal of improving sensitivity to such therapies by combining them with perturbations in specific components of the PCD network.
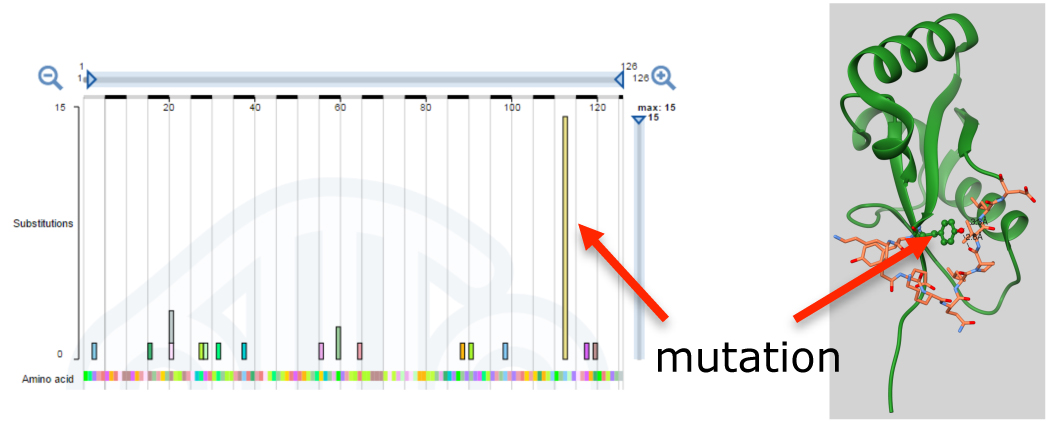
- Using bioinformatics tools, we have identified somatic mutations in autophagy genes that affect their autophagy functions and are linked to melanoma and carcinoma.