Most proteins function within a narrow range of salt concentration. However, in a project carried out in close collaboration with Prof. Ada Zamir, Dept of Biomolecular Sciences, studies of halotolerant Dunaliella salina revealed a new class of proteins - halotolerant proteins - that can function in the entire range of salinity (0 to 4 M NaCl). In order to get an insight into molecular details of how do these proteins take up the challenge to function in the varying salt concentrations, we have determined the first crystal structure of a halotolerant protein - a 30kDa Carbonic anhydrase.
The crystal structure coupled with the series of site-directed mutants will aid in for the better understanding of: what is the unique nature of protein - solvent and weak interactions for these class of proteins? What is the structural principle that confers the additional stability of halotolerance. How do halotolerant enzyme protect themselves from anionic/ cationic inhibition? Could this understanding have biotechnological applications such as engineering the enzymes to function in special solvents for industrial or environmental use?
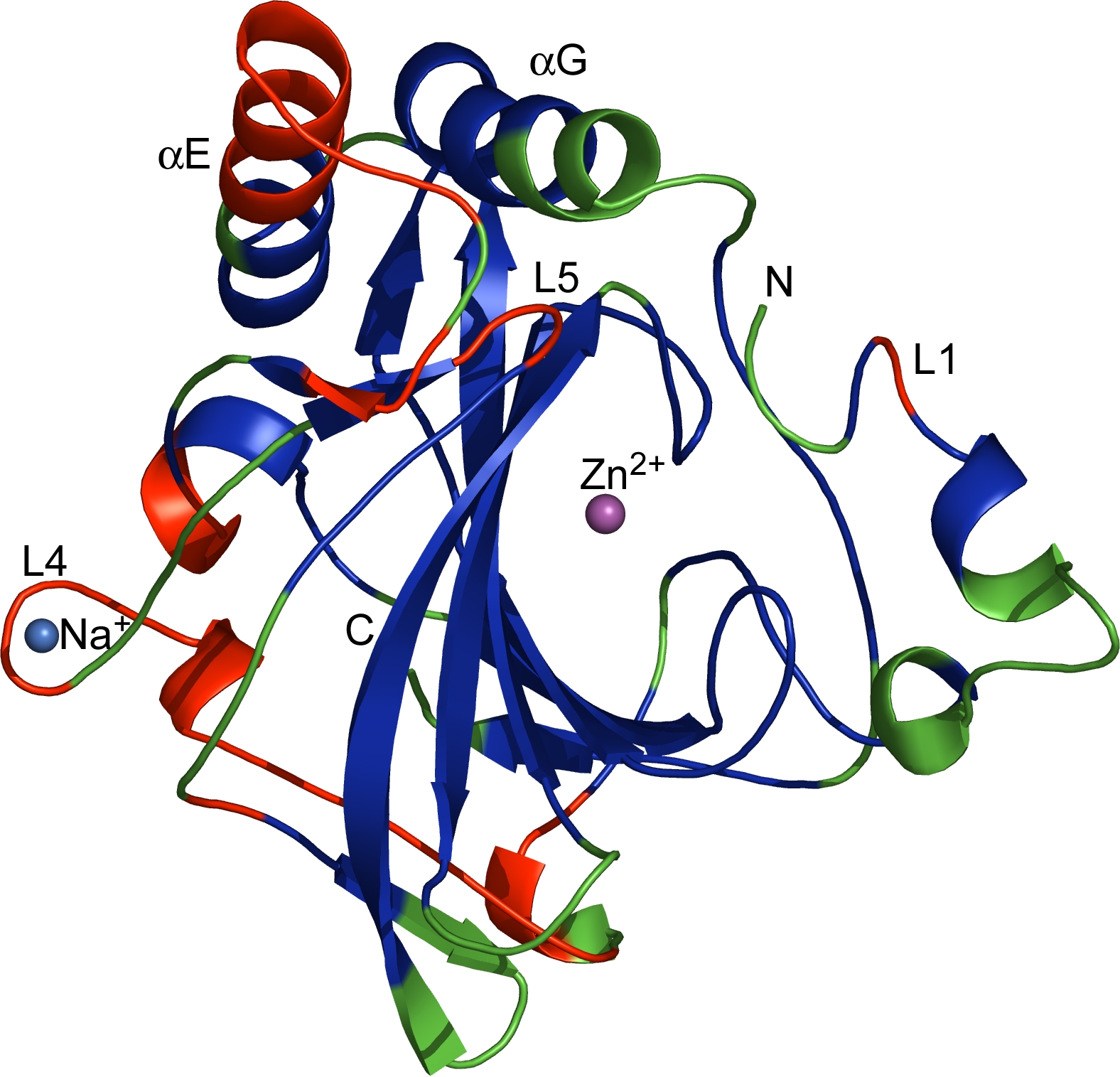
The regions corresponding to CRs (blue), VRs (green) and VCRs (red), as defined in the text, are mapped onto the dCA II structure. Marked are the catalytic Zn2+, insertions and deletions in VCRs including L1 (the Zn binding loop), L4 (the Na-binding loop) and L5 as well as the two extended -helices (E and G). N- and C-termini are indicated.


