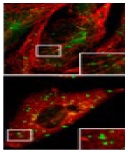
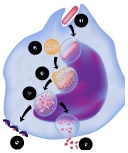
Understanding the process of intracellular protein transport, whereby protein-carrying vesicles move across and between membranes, is of fundamental importance in cell physiology and medicine. Autophagy (literally self-eating) is a unique membrane trafficking process essential for the degradation and recycling of excess and/or defective macromolecules and organelles in eukaryotes. This pathway is activated under environmental stress conditions as well as during certain developmental stages and has been linked to programmed cell death, cancer, pathogen infection, and degradation of ubiquitinated protein aggregates formed in many pathological conditions. The main goal of our studies is to understand the molecular mechanism of autophagy and characterize its exact role in health and disease employing mammalian, and yeast systems.
Zvulun (Zevi) Elazar
Harold Korda Professor at the Department of Biological Chemistry of the Weizmann Institute of Science in Rehovot, Israel.
Zvulun studies different mechanistic aspects of autophagy in yeast and mammalian systems. He is particularly interested in the role of the Atg8s, unique ubiquitin-like proteins that are part of the core autophagic machinery. This family facilitates both autophagosome formation and selective cargo recruitment into these organelles. The current focus of his lab is on the identification of factors that regulate these processes and are defective in pathophysiological conditions such as neurodegeneration and cancer.