One fish, two fish
Briefs
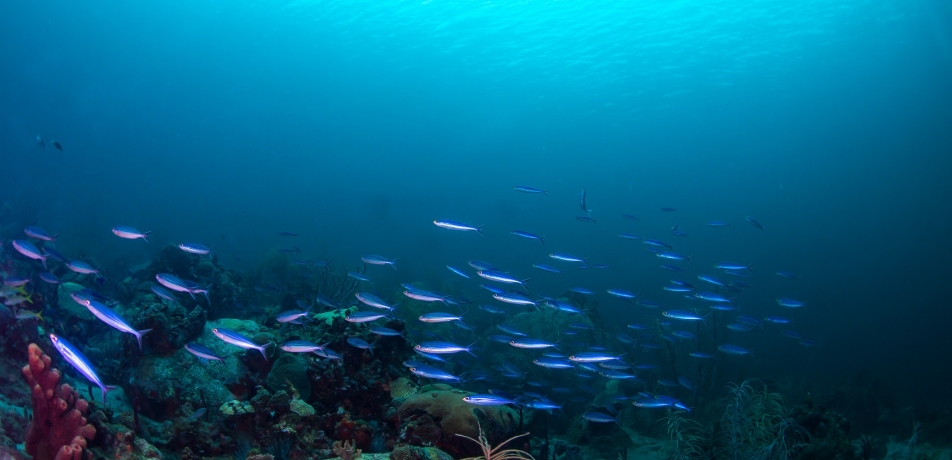

To marine biologists who monitor the gulf between Eilat and Aqaba in the Red Sea, population counts of the tiny larvae that develop into mature fish are an important indicator of ecological health. But with more than 500 fish species known to frequent the gulf’s coral reefs, and with the dispersal of fish larvae governed by chaotic underwater currents, identifying and tracking individual species has been nearly impossible—until now.
Prof. Rotem Sorek of the Department of Molecular Genetics has developed a new “barcoding” technique that allows scientists to monitor not only which larvae species are present in the Gulf of Aqaba, but how many of each, at what time of year, and at which depths. Collaborating with marine biologists from Tel Aviv University and Ben-Gurion University whose previous research involved inspecting larvae under a microscope and counting spines, Prof. Sorek’s new technique was based on the sequencing of larval genomes. This work was conducted at the Stephen and Nancy Grand Israel National Center for Personalized Medicine on campus.
To gather the relevant data, the scientists took up diving in the Gulf, clipping the fins of adult fish for DNA analysis. Prof. Sorek and his team then used a “metagenomics” technique to match this adult genetic material to larvae, ultimately achieving barcodes for around 80% of the fish species.
“Although the initial process was a bit arduous, we now have an excellent tool for monitoring the health of the reef ecosystem, and other reef researchers may follow suit,” Prof. Sorek says.
Prof. Rotem Sorek is supported by the Abisch Frenkel Foundation for the Promotion of Life Sciences, The Y. Leon Benoziyo Institute for Molecular Medicine, the European Research Council, the Leona M. and Harry B. Helmsley Charitable Trust , the Dana and Yossie Hollander Center for Structural Proteomics, Martin Kushner Schnur, and the David and Fela Shapell Family Foundation G-INCPM Fund for Preclinical Studies.