Genomes of living organisms are comprised of very long DNA molecules. A fundamental question is by what mechanisms are specific loci along these genomes found, with high efficiency and at relevant physiological times. We address this question in the case of horizontal gene transfer processes such as conjugation and viral transduction, which result in the rapid acquisition of new traits in bacteria.
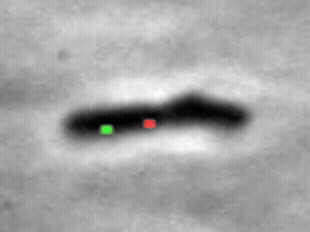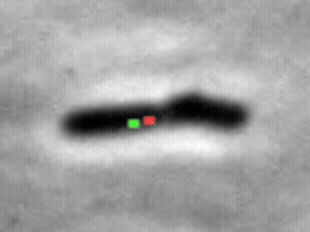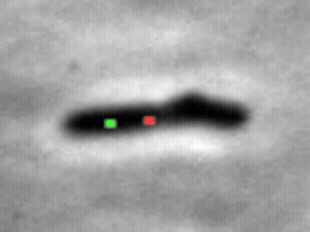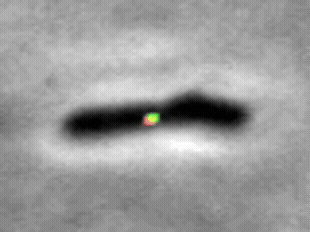
We study the import of integrating conjugating elements (ICEs, in green) by B. subtilis recipient cells, and their search for their unique integration sites along the host chromosome (red).

We also show that chromosomal sites in B. subtilis cells move sub-diffusely, driven by the out-of-equilibrium nature of the viscoelastic, crowded intracellular environment. This in vivo motion, which is asymmetric, is due to chromosomal architecture, as supported by molecular dynamics simulations of chromosomal loops in bottlebrush chromosomes of bacteria.

Previously, we also studied the infection of E. coli cells by bacteriophage lambda, whose DNA integrates at a unique site into the bacterial genome, when following the lysogenic pathway. To shed light on the mechanisms by which lambda DNA finds its unique integration site, we follow in real time individual lambda DNAs and their integration site within live cells using fluorescent markers, until lysogeny is established, revealing the dynamics of the search process.