Our ultimate goal is to harness our knowledge of the PCD network and its governing principles in order to more fully understand the etiology of disease in which cell death and related pathways are misregulated, such as cancer, and ultimately, to develop PCD-based therapies.
The PCD Interactome
We developed a protein-protein interaction (PPI) platform using the Protein-fragment Complementation Assay (PCA) to establish the protein interactome of the cell death network. The PCA assay is based on the fusion of two individual proteins to either half of a luciferase reporter. When they interact, the two halves of the luciferase protein are brought into proximity and fold into the active enzyme. Thus luciferase activity is a quantitative read-out of the strength of interaction between the two proteins. Using a comprehensive PCA library of PCD proteins fused to either Luc fragment, we generated a detailed PPI landscape of the apoptosis and autophagy modules and points-of-interface between them. Some of our insights included identifying previously unknown interactions within modules, such as WIPI2/Ulk1 and DAPK2/14-3-3, and PPIs between proteins within different modules, serving as points-of-entry to novel pathways that regulate inter-modular cross-talk.

Schematic describing PCA platform to identify PPIs within the PCD network. From Gilad, et al., Cell Rep. 2014. 8:909-21.
More recently, we have taken the PPI interactome a step forward, as the basis for novel drug discovery. Critical PPIs within the network provide more specific targets for intervention than their component protein nodes, which can individually function in multiple pathways. Moreover, the binding interface provides an optimal target for small molecules that can block the interaction, and thus interfere with a critical step within a particular pathway. With this in mind, we chose the Atg12-Atg3 PPI to be targeted for the purpose of developing efficient and specific autophagy inhibitors, which are currently lacking in the field. We developed a small molecule inhibitor pipeline based on the Atg12-Atg3 PCA, which has yielded several interesting potential drug candidates. These have been applied to various cell models in which autophagy contributes to disease etiology, such as autophagy-addicted cancer cells, and autophagy-dependent inflammation, as a first step in determining the benefit of specific autophagy inhibition to disease treatment.
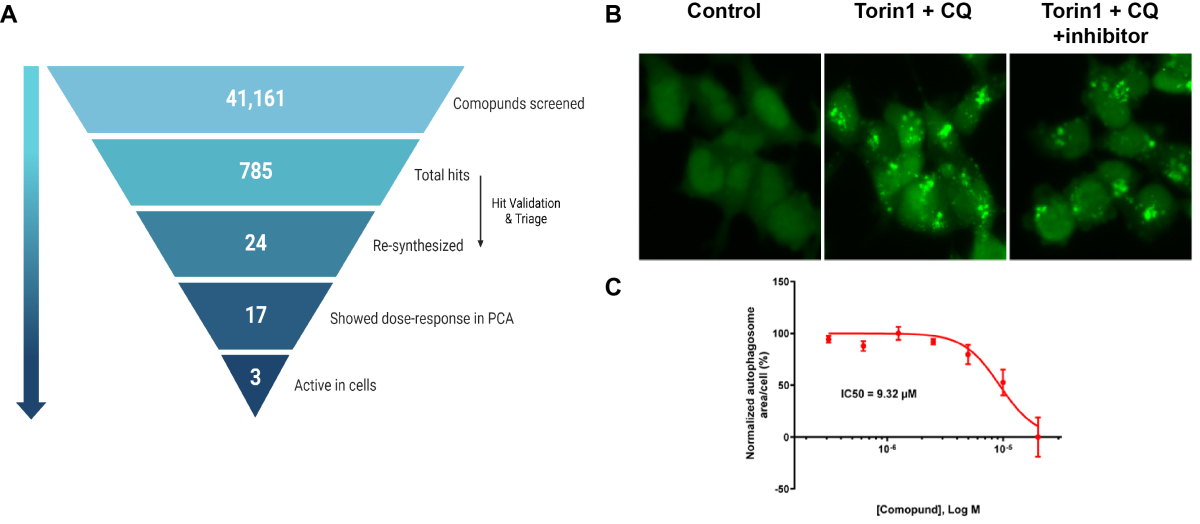
A. Summary of PPI-based drug screen. B. Effects of one inhibitor on autophagosome formation (green puncta, GFP-LC3) following induction of autophagy with mTOR inhibitor Torin1 and lysosomal inhibitor chloroquine (CQ). C. Dose response curve for inhibitor based on assay shown in B. From Nuta, et al., 2023, Autohagy, 19:2372-2385.
A second PPI target that we are currently exploring is the DAPK2 homodimer. DAPK2 (see Appendix) has been shown to be a tumor suppressor, commonly attributed to its ability to induce cell death, block anchorage independent growth by promoting anoikis and/or prevent chemoresistance. Its expression is down-regulated in several cancers, and has also been correlated with poor prognosis. Activation of the residual DAPK2 can therefore be a means to restore DAPK2 function and include cell death, as a novel targeted cancer therapy. The resolved crystal structure of DAPK2 shows that it forms homodimers in a closed auto-inhibited conformation that blocks substrate access but allows ATP binding to the nucleotide binding site. Disassembly of the DAPK2 homodimer enables Ca2+/CaM binding and kinase activation to allow substrate phosphorylation. The interaction surface involves a basic loop motif of one monomer that inserts into a groove between two helices in the opposing monomer, and serves as a promising target for perturbation by small molecules. Our current goal is to use the PCA-based drug screening strategy to identify PPI inhibitory compounds that target the dimerization interface of DAPK2, causing homodimer disassembly and kinase activation.

Ribbon diagram depicting structure of DAPK2 dimer with basic loop in yellow.