Yeast-based
To answer some of the questions raised above. We are currently using yeast-based screens to identify elements involved in gating specificity just preceding channel pore openings, which were otherwise impossible to detect by conventional biochemical or structure-functional approaches. We also employ various molecular techniques to understand channel function at the single molecule level mainly by using patch clamp single channel recordings.
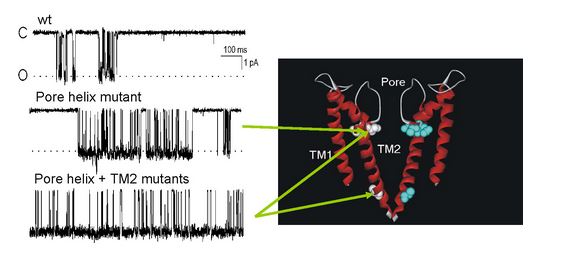
Fig 3: Single channel recordings traces (right) of mutants shown in the structure model of KcsA (left).
Conformational dynamics and, signaling specificity of channel gating and intramolecular protein rearrangements are being investigated using fluorescence resonance energy transfer (FRET) combined with advanced microscopy techniques. These studies also include the investigation of G protein modulation of neuronal calcium channels and ionic homeostasis in the mitochondria.
Fluorescence
We are currently developing mouse lines to study, both neurological manifestation of Down’s syndrome, the involvement of the channel in the immune system and normal ion channel functional plasticity using fluorescence techniques (Fig. 4). Metabolic regulation and subcellular localization of the channels are being investigated at two levels, one using conventional biochemical and electrophysiological approaches and second by the mass production of channel domains for structural and proteomic studies. Finally, a more general fluorescent based approach is being developed to detect G protein associated signaling in living cells for the generation of both nanosensors for GPCR activity and for orphan GPCRs drug discovery approaches.

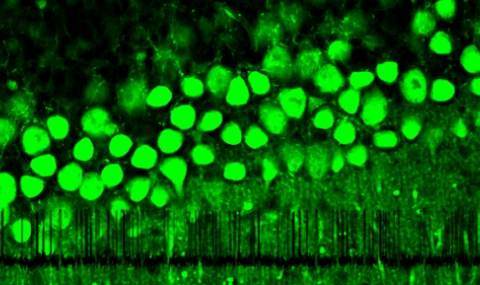
Fig 4: Brain from a wt and knockin mice where the GIRK1 gene was replaced with a wt gene fused with yellow fluorescent protein (YFP). Left panel, a fluorescent image (only the brain from the knockin mouse is fluorescing- right brain). Right panel, the same brains under normal illumination from the bottom.