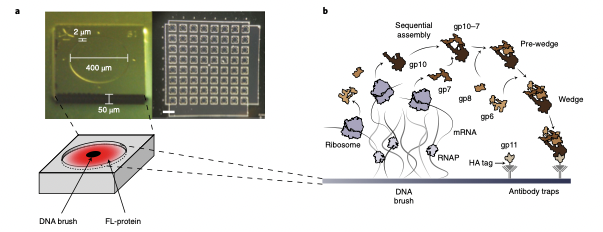
The assembly of protein machines in cells is precise, rapid, and coupled to protein synthesis with regulation in space and time. The assembly of natural and synthetic nanomachines could be similarly controlled by genetic programming outside the cell. We created quasi-two-dimensional (2D) silicon compartments that enable programming of protein assembly lines by local synthesis from surface-immobilized DNA brushes. Using this platform, we studied the autonomous synthesis and assembly of a structural complex from a bacteriophage and a bacterial RNA-synthesizing machine. Local synthesis and surface capture of complexes provided high assembly yield and sensitive detection of spatially resolved assembly intermediates, with the 3D geometry of the compartment and the 2D pattern of brushes dictating the yield and mode of assembly steps. Localized synthesis of proteins in a single gene brush enhances their interactions, and displacement of their genes in separated brushes leads to step-by-step surface assembly. This methodology enables spatial regulation of protein synthesis, and deciphering, reconstruction and design of biological machine assembly lines.