Identifying heterogeneity in clinical samples
Complicated bacterial infections are often difficult to treat and frequently recur, as in urinary tract infections. One reason is the presence of bacterial subpopulations that survive therapy by occupying distinct physiological states. We aim to identify and characterize these co-existing bacterial states directly in clinical samples by combining Microcolony-seq with experimental and computational tools.

Using this approach in a Enteropathogenic E. coli (EPEC) isolate, in a Uropathogenic E. coli (UPEC) urine infection and in a Staphylococcus aureus (S. aureus) bloodstream infection, we have identified distinct bacterial subpopulations co-existing within individual patients. These subpopulations are genetically identical yet phenotypically diverse, exhibiting differences in metabolic activity and, importantly, in their virulence states.
Building on this work, we aim to extend our analyses to additional patients and to bacterial infections from multiple clinical sites. Our goal is to broaden our understanding of how co-existing bacterial subpopulations arise and contribute to recurrence during infection.
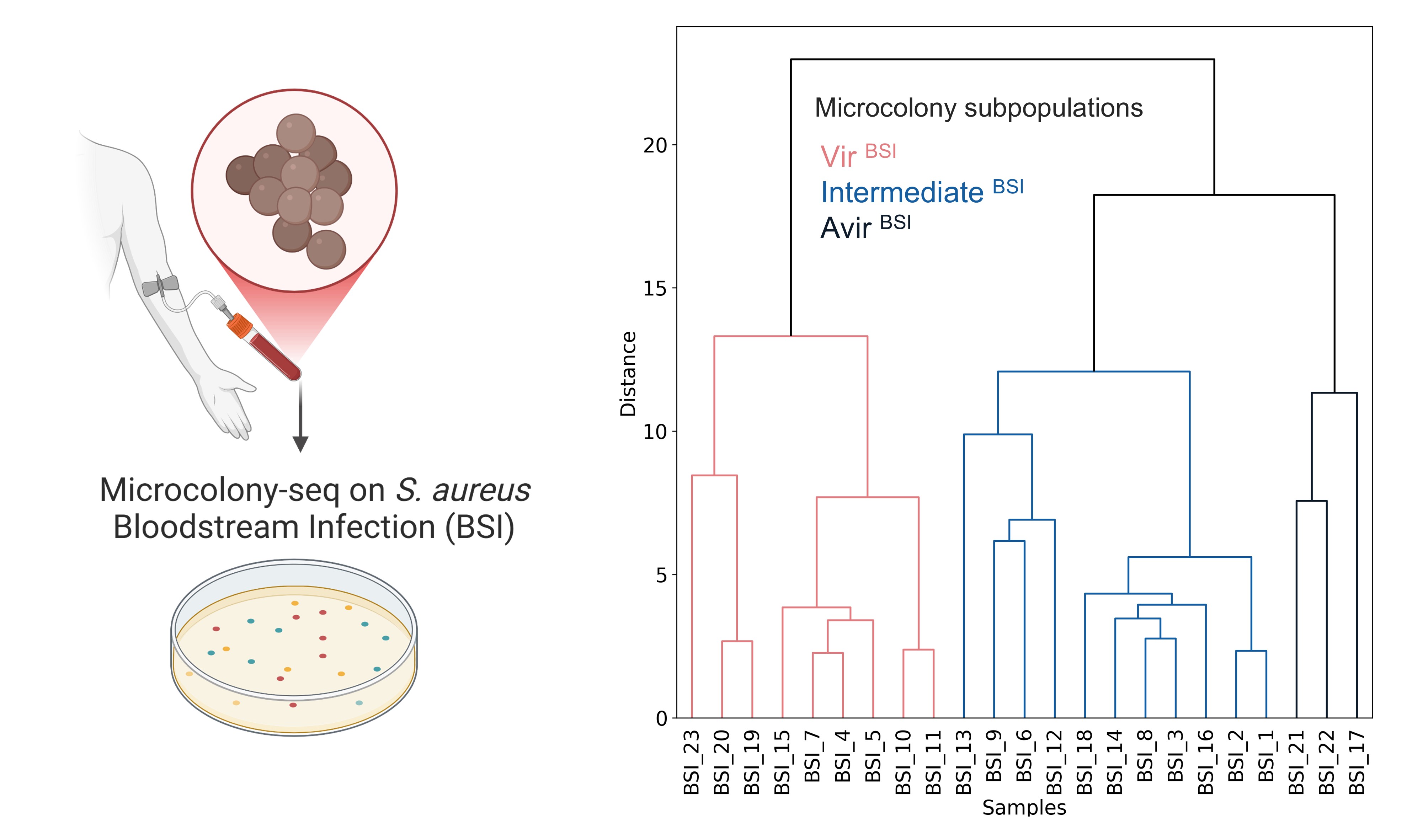
Faigenbaum-Romm et al., "Uncovering phenotypic inheritance from single cells with Microcolony-seq". Cell, 2025.
