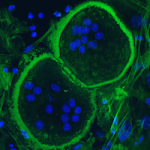
Phosphorylation of tyrosine residues in proteins is a major mechanism for regulating their structure and function. Phosphorylation is reversible and is regulated by the generically opposing activities of tyrosine kinases and tyrosine phosphatases (PTPs).
Over 100 PTP genes are known. The products of these genes exhibit substrate specificity that ranges from exclusively phospho-tyrosine through phospho-serine and –threonine and up to lipids and RNA. These enzymes are classified as PTPs since their catalytic mechanism is similar and is characteristic of the tyrosine-specific, so-called “classical” PTP family, which was the first to be discovered by Nicholas Tonks and Edmond Fischer in 1988.
A very large number of studies has established that dysregulation of protein phosphorylation can lead to serious disease, including cancer, metabolic syndrome, neural dysfunction, and others. Molecules that regulate these processes are therefore critical modulators of cellular function, as has been shown by increasing development and use of modulators of tyrosine kinase activity as drugs for treating disease. Tyrosine phosphatases play roles of similar importance in these processes, and their specificity in vivo is high. Since they are entirely distinct from tyrosine kinases, study of PTPs and their molecular roles in physiological processes can shed new light on the molecular details of these processes and suggest novel ways to intervene in them for therapeutic gain.