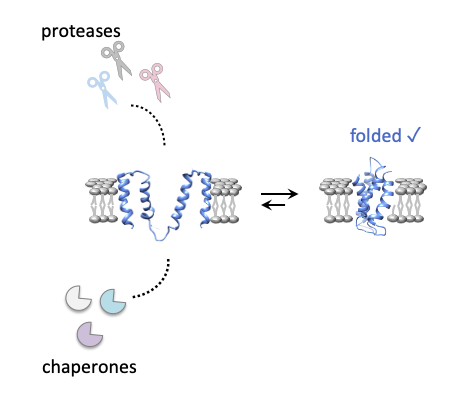
The vast majority of proteins in nature need to fold to perform their function. Folding is essential for life, but it is an incredibly complex process that quite frequently fails, leading to an enormous diversity of diseases. Evolution has equipped both the protein sequence and the cellular machinery with sophisticated mechanisms to promote folding and to aid the cell getting rid of misfolded proteins. Membrane proteins reside within the hydrophobic membrane and it is clear that they undergo folding and quality control by mechanisms that are quite distinct from globular proteins – but these mechanisms still remain largely elusive.
Membrane protein homeostasis seems to be an Achilles’ heel for eukaryotic and prokaryotic cells alike. In prokaryotes, membrane protein damage is a primary factor in the toxicity of antibiotics. In eukaryotes, some membrane proteins are apparently so hard to produce, that well over 50% are immediately degraded, with many destabilizing mutations causing disease. How do mutations destabilize membrane proteins? What determinants attract cellular quality control and how are they recognized? These questions remain open even for some of the best characterized model systems. Studies in eukaryotes are especially complicated by trafficking along the secretory compartments with its multiple quality control checkpoints. Bacteria, on the other hand, offer a simpler environment to dissect some of the fundamental biology. We approach this topic by a analyzing the relationship between protein sequence, folding state and quality control factors. We use a combination of low and high throughput quantitative techniques, including biochemistry, genetics and genetic screening, biophysics, flow cytometry and in-vivo stability assays.